Systems-Biology using GoogleGears: my notebook
Google gears is an open source browser extension that enables web applications to provide offline functionality. The data are stored locally in a fully-searchable relational database using the sqlite engine.
My Biological Network is a tool I created as a test to play with Google gears: it is used to build a network of protein-protein interactions. It uses Google Gears to record your entries on the local disk, so Gears needs to be installed on your computer. Programming with gears with JAVASCRIPT is really cool as you don't have to implement the storage of the data on the server side and you're using some standard SQL statements to handle the data.
Screenshots
Tutorial
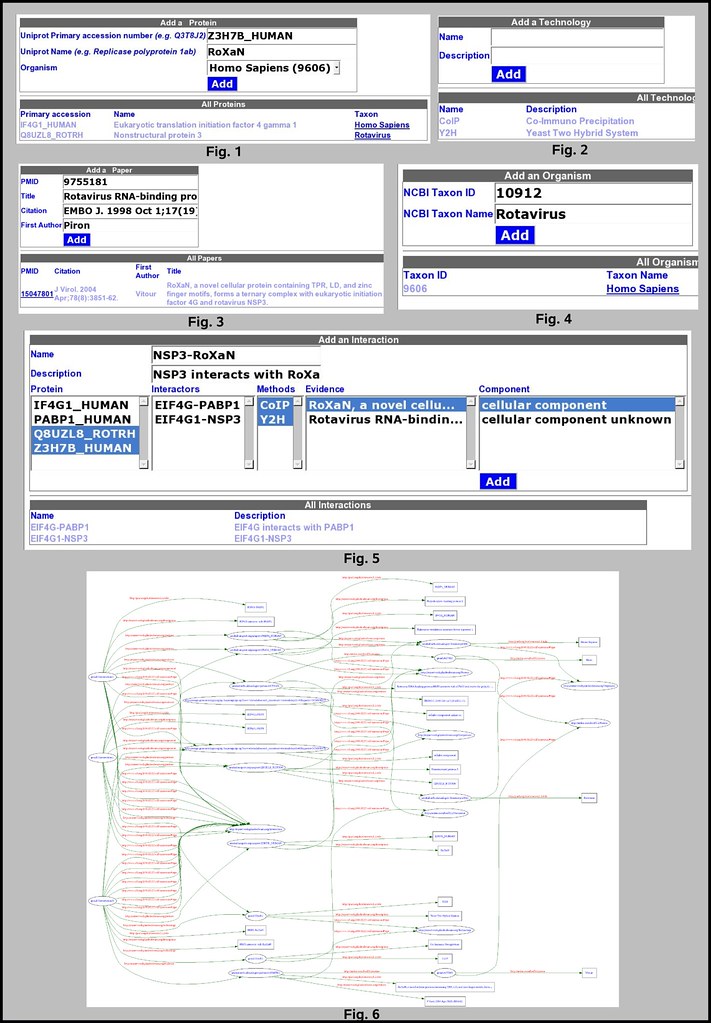
Open the tab Organism (fig. 4): add one or more organism. (Homo Sapiens already inserted by default)
Open the tab Protein (fig. 1): add one or more protein.
Open the tab Paper (fig. 3): add one or more article that will be used as an evidence for an interaction.
Open the tab Technology (fig. 2): add one or more technology that was used to characterize an interaction.
Open the tab Component: add one or more cellular component using Gene Ontology (GO:0005575 \"cellular component\" was inserted by default)
Open the tab Interaction (fig. 5):
- Name and describe this interaction
- Select one or more protein and/or one or more previously defined proteic complex. You Cannot describe self interactions with this tool.
- (optional) choose one or more paper/technology/component...
Open the RDF table (fig. 6): I choose to display the content of the database using RDF. Such format can then be validated and visualized using the W3C RDF validator, or transformed using XSLT, etc.... I also used the life science identifier (LSID) as an URI for my resources.
On my computer, the database is stored in /env/islande/home/lindenb/.mozilla/firefox/<profile-id>/Google Gears for Firefox/islande/<host>/mynetwork#database. The database can be manualy accessed using sqlite3:
sqlite3 mynetwork#database
SQLite version 3.4.0
Enter '.help' for instructions
sqlite> .tables
component interactionhash paper technology
interaction organism prote
sqlite> .schema organism
CREATE TABLE organism(id integer primary key ,name varchar(50) not null unique);
sqlite> select * from organism;
9606|Homo Sapiens
sqlite>
Internals
We the page is loaded, we check that gears was installed
if (!window.google || !google.gears) {
debug("NOTE: You must install Google Gears first.")
We then create the database if does not exist. The file is created in firefox in ${HOME}/.mozilla/firefox/<profile-id>/Google Gears for Firefox/<server>/mynetwork#database
connection = google.gears.factory.create("beta.database","1.0");I create the tables just by invoking some standards SQL 'CREATE TABLE' statements. I also insert some default values (e.g. human organism)
connection.execute("create table if not exists organism(id integer primary key ,name varchar(50) not null unique)");
connection.execute("insert or ignore into organism(id,name) values(9606,\"Homo Sapiens\")");
connection.execute("create table if not exists protein(id integerprimary key autoincrement,name varchar(50) not null,taxId int not null,acn varchar(50) not null unique)");
connection.execute("create table if not exists paper(pmid integerprimary key ,title varchar(255) not null,citation varchar(255) not null,firstAuthor varchar(50) not null)");
connection.execute("create table if not exists component(id integer primary key autoincrement,go varchar(50) not null unique, name varchar(50) not null unique)");
connection.execute("insert or ignore into component(go,name) values(\"GO:0005575\",\"cellular component\")");
connection.execute("insert or ignore into component(go,name) values(\"GO:0008372\",\"cellular component unknown\")");
connection.execute("create table if not exists technology(id integer primary key autoincrement,name varchar(50) not null unique, description varchar(255) not null)");
connection.execute("insert or ignore into technology(name,description) values(\"Y2H\",\"Yeast Two Hybrid System\")");
connection.execute("insert or ignore into technology(name,description) values(\"CoIP\",\"Co-Immuno Precipitation\")");
connection.execute("create table if not exists interaction(id integer primary key autoincrement, name varchar(50) not null unique,description varchar(255) not null)");
connection.execute("create table if not exists interactionhash(id integer primary key autoincrement,LINK_interaction int ,type varchar(20) not null,child int not null)");
When a data is about to be inserted we check all the fields and we insert them using SQL: INSERT INTO
var id= getById("organism-input-id");
if(!isInteger(id.value))
{
debug("TaxId not a Number");
return;
}
var name=getById("organism-input-name");
if(trim(name.value).length==0)
{
debug("Taxon Name empty");
return;
}
try
{
connection.execute("insert into organism(id,name) values("+sqlescape(trim(id.value))+","+sqlquote(trim(name.value))+")");
id.value="";
name.value="";
}
catch(err)
{
debug(err.message);
return;
}
a simple SELECT is used to retrieve the data and insert them in a HTML table
var rs= connection.execute("select id,name from organism order by name");
while (rs.isValidRow())
{
var tr= ce("tr");
table.appendChild(tr);
var td= ce("td");
tr.appendChild(td);
td.appendChild(ct(rs.field(0)));
td= ce("td");
tr.appendChild(td);
var a= ce("a");
a.setAttribute("title","Open in NCBI");
a.setAttribute("target","tax"+rs.field(0));
a.setAttribute("href","http://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?id="+rs.field(0));
td.appendChild(a);
a.appendChild(ct(rs.field(1)));
rs.next();
}
rs.close();
That's it !
Pierre
updated 2010-08-12: source code
<html xmlns="http://www.w3.org/1999/xhtml">
<head>
<script type="text/javascript" src="gears_init.js"></script>
<script type="text/javascript" src="network.js"></script>
<link rel="stylesheet" type="text/css" href="./network.css" />
<title>My Biological Network</title>
</head>
<body onload="init()">
<h1>My Biological Network</h1>
<p>Pierre Lindenbaum PhD <a href="mailto:plindenbaum@yahoo.fr">plindenbaum@yahoo.fr</a><br/><a href="http://plindenbaum.blogspot.com">http://plindenbaum.blogspot.com</a><br/><address>Bioinformatics department<br/><a href="http://www.integragen.com">Integragen S.A.</a><br/>Evry, France</address></p>
<p/>
<div>
<button onclick="javascript:showCard('home-pane');">Home</button>
<button onclick="showOrganismPane()">Organisms</button>
<button onclick="showProteinPane()">Proteins</button>
<button onclick="showPaperPane()">Papers</button>
<button onclick="showTechnologyPane()">Technology</button>
<button onclick="showComponentPane()">Component</button>
<button onclick="showInteractionPane()">Interactions</button>
<button onclick="showRDFPane()">RDF</button>
</div>
<div style="color:red;" id="stderr"></div>
<p/>
<!-- ====================================== ORGANISM ====================================== -->
<div style="display:none;" id="organism-pane">
<table>
<caption>Add an Organism</caption>
<tr><th>NCBI Taxon ID <i>(e.g. 10912)</i></th><td><input id="organism-input-id" length="10"/></td></tr>
<tr><th>NCBI Taxon Name <i>(e.g. Rotavirus)</i></th><td><input id="organism-input-name" length="10"/></td></tr>
<tr><th/><td><button onclick="addOrganism()">Add</button></td></tr>
</table>
<hr/>
<table width="80%">
<caption>All Organisms</caption>
<thead>
<tr><th>Taxon ID</th><th>Taxon Name</th></tr></tr>
</thead>
<tbody id="organism-table">
</tbody>
</table>
</div>
<!-- ====================================== COMPONENT ====================================== -->
<div style="display:none;" id="component-pane">
<table>
<caption>Add a Component</caption>
<tr><th>Name</th><td><input id="component-input-name" length="10"/></td></tr>
<tr><th>GO</th><td><input id="component-input-go" length="10"/></td></tr>
<tr><th/><td><button onclick="addComponent()">Add</button></td></tr>
</table>
<hr/>
<table width="80%">
<caption>All Components</caption>
<thead>
<tr><th>Name</th><th>GO</th></tr>
</thead>
<tbody id="component-table">
</tbody>
</table>
</div>
<!-- ====================================== TECHNOLOGY ====================================== -->
<div style="display:none;" id="technology-pane">
<table>
<caption>Add a Technology</caption>
<tr><th>Name</th><td><input id="technology-input-name" length="50"/></td></tr>
<tr><th>Description</th><td><input id="technology-input-desc" length="50"/></td></tr>
<tr><th/><td><button onclick="addTechnology()">Add</button></td></tr>
</table>
<hr/>
<table width="80%">
<caption>All Technologies</caption>
<thead>
<tr><th>Name</th><th>Description</th></tr></tr>
</thead>
<tbody id="technology-table">
</tbody>
</table>
</div>
<!-- ====================================== PROTEIN ====================================== -->
<div style="display:none;" id="protein-pane">
<table>
<caption>Add a Protein</caption>
<tr><th>Uniprot accession number <i>(e.g. Q3T8J2)</i></th><td><input id="protein-input-acn" length="10"/></td></tr>
<tr><th>Uniprot Name <i>(e.g. Replicase polyprotein 1ab)</i></th><td><input id="protein-input-name" length="10"/></td></tr>
<tr><th>Organism</th><td><select id="protein-input-taxon" length="10"><option>A</option></select></td></tr>
<tr><th/><td><button onclick="addProtein()">Add</button></td></tr>
</table>
<hr/>
<table width="80%">
<caption>All Proteins</caption>
<thead>
<tr><th>Primary accession</th><th>Name</th><th>Taxon</th></tr></tr>
</thead>
<tbody id="protein-table">
</tbody>
</table>
</div>
<!-- ====================================== PAPER ====================================== -->
<div style="display:none;" id="paper-pane">
<table>
<caption>Add a Paper</caption>
<tr><th>PMID</th><td><input id="paper-input-pmid" length="10"/></td></tr>
<tr><th>Title</th><td><input id="paper-input-title" length="50"/></td></tr>
<tr><th>Citation</th><td><input id="paper-input-citation" length="50"/></td></tr>
<tr><th>First Author</th><td><input id="paper-input-author" length="50"/></td></tr>
<tr><th/><td><button onclick="addPaper()">Add</button></td></tr>
</table>
<hr/>
<table width="80%">
<caption>All Papers</caption>
<thead>
<tr><th>PMID</th><th>Citation</th><th>First Author</th><th>Title</th></tr></tr>
</thead>
<tbody id="paper-table">
</tbody>
</table>
</div>
<!-- ====================================== INTERACTION ====================================== -->
<div style="display:none;" id="interaction-pane">
<table>
<caption>Add an Interaction</caption>
<tr><th>Name</th><td colspan="4"><input id="interaction-input-name" length="50"/></td></tr>
<tr><th>Description</th><td colspan="4"><input id="interaction-input-desc" length="50"/></td></tr>
<tr>
<th>Protein</th>
<th>Interactors</th>
<th>Methods</th>
<th>Evidences</th>
<th>Components</th></tr>
<tr>
<td><select id="interactors-input-proteins" size="5" multiple="true"/></td>
<td><select id="interactors-input-interactors" size="5" multiple="true"></td>
<td><select id="interactors-input-technologies" size="5" multiple="true"></td>
<td><select id="interactors-input-evidences" size="5" multiple="true"></td>
<td><select id="interactors-input-components" size="5" multiple="true"></td>
</tr>
<tr><th colspan="4"/><td><button onclick="addInteraction()">Add</button></td></tr>
</table>
<hr/>
<table width="80%">
<caption>All Interactions</caption>
<thead>
<tr><th>Name</th><th>Description</th></tr></tr>
</thead>
<tbody id="interaction-table">
</tbody>
</table>
</div>
<!-- ====================================== RDF ====================================== -->
<div style="display:none;" id="rdf-pane">
<h2>RDF Pane</h2>
<textarea wrap="off" id="rdf-area" rows="20" cols="80"></textarea>
</div>
<!-- ====================================== HOME ====================================== -->
<div style="display:none;" id="home-pane">
<h3>About My Biological Network</h3>
<p><a href="http://gears.google.com/">Google gears</a> is an open source browser extension that enables web applications to provide offline functionality. The data are stored locally in a fully-searchable relational database using the <a href="http://www.sqlite.org/">sqlite engine</a>.</p>
<p><b>My Biological Network</b> is a tool I created as a test to play with Google gears: it is used to build a network of protein-protein interactions. It uses Google Gears to record your entries on the <u>local disk</u>, so Gears needs to be installed on your computer. </p>
<p>
Open the tab <b>Organism</b>: add one or more organism. (Homo Sapiens already inserted by default)<br/>
Open the tab <b>Protein</b>: add one or more protein.<br/>
Open the tab <b>Paper</b>: add one or more article that will be used as an evidence for an interaction.<br/>
Open the tab <b>Technology</b>: add one or more technology that was used to characterize an interaction.<br/>
Open the tab <b>Component</b>: add one or more cellular component using Gene Ontology (GO:0005575 \"cellular component\" was inserted by default)<br/>
Open the tab <b>Interaction</b>:<ul>
<li>Name and describe this interaction</li>
<li>Select one or more protein and/or one or more previously defined proteic complex. You <i>Cannot</i> describe self interactions with this tool.<li>
<li>(optional) choose one or more paper/technology/component...</li>
</ul><br/>
Open the <b>RDF table</b>: I choose to display the content of the database using <a href="http://www.w3.org/RDF/">RDF</a>. Such format can then be validated and visualized using the <a href="http://www.w3.org/RDF/Validator/">W3C RDF validator</a>, or transformed using <a href="http://www.w3.org/TR/xslt">XSLT</a>, etc.... I also used the <a href="http://lsid.sourceforge.net/">life science identifier (LSID)</a> as an URI for my resources.<br/>
</p>
<p>On my computer, the database is stored in <code>$HOME/.mozilla/firefox/<profile-id>/Google Gears for Firefox/<host>/mynetwork#database</code>. The database can be manualy accessed using <a href="http://www.sqlite.org/">sqlite3</a>:<pre style='color:black;border:1pt solid;background:lightgray;'>sqlite3 mynetwork#database
SQLite version 3.4.0
Enter '.help' for instructions
sqlite> .tables
component interactionhash paper technology
interaction organism prote
sqlite> .schema organism
CREATE TABLE organism(id integer primary key ,name varchar(50) not null unique);
sqlite> select * from organism;
9606|Homo Sapiens
sqlite></pre>
</p>
</div>
<!-- google analytics -->
<script src="http://www.google-analytics.com/urchin.js"
type="text/javascript">
</script>
<script type="text/javascript">
_uacct = "XXXXXX";
urchinTracker();
</script>
<!-- google analytics -->
</body>
</html>
3 comments:
Pierre, you rock.
I couldn't get anything to appear in the interactions list, though...
I just tested it again: I worked fine for me (?)
Have you tried with SBML ?
Sincerely
Mr BioS i5i :)
Post a Comment